Introduction
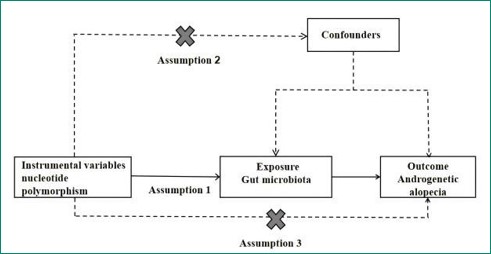
Androgenetic Alopecia (AGA) is one of the most prevalent types of hair loss in clinical practice, characterized by a significant genetic predisposition and a close correlation with androgen levels in the human body [1]. In China, over 250 million individuals are facing issues related to hair loss, with the onset age of this condition showing a declining trend, which has had a substantial impact on personal image [2]. AGA primarily results from excessive secretion of androgens in the scalp or heightened sensitivity to these hormones. The hallmark features of this condition include miniaturization of hair follicle cells in the frontal and crown areas as well as a reduction in the growth phase duration of dermal cells. This leads to an increasing density of damaged hair follicles and subsequently causes gradual hair thinning [3,4]. The Gut Microbiota (GM) refers to the vast collection of microorganisms residing within the gastrointestinal tract. These microbes play crucial roles by absorbing nutrients that stimulate intestinal cells and participating in immune responses, thereby influencing the host’s immune system. With ongoing research into the Gut-Skin Axis, it has become evident that GM plays an important role in maintaining systemic immune homeostasis [5]. Due to bidirectional communication between GM and skin health, various pathways such as immune modulation and transfer of strains or metabolic products can affect skin health [6-8]. MR is a research methodology that utilizes randomly assigned genetic variations within populations as instrumental variables. By establishing associations between these instrumental variables, exposure factors, and outcomes, MR aims to reveal potential causal relationships between exposure factors and outcomes [9]. Given that genetic variations are randomly allocated during meiosis-once alleles are formed, they remain unaffected by subsequent confounding factors- MR can simulate effects akin to those observed in randomized controlled trials [10,11]. This study employs MR methods to analyze causal links between GM and AGA with an aim to provide reference for future research (Figure 1).
Histological evaluation found a lymphocyte infiltrate consisting
of a prevalence of small-sized T lymphocytes (CD3+ and CD5+)
with a mixture of CD4+ and CD8 + lymphocytes with a small count
of B+ lymphocytes (CD 20+). In the samples there were no elements referable to chronic IBD and no evidence of Kaposi’s sarcoma. At immunohistochemistry and citofluorimetry no signs of
monoclonality or images suggestive for lymphoma were seen.
Therefore, after ruling out opportunistic infections, on April
2021 he started Highly Active Antiretroviral Therapy (HAART)
based on bictegravir, emtricitabine and tenofovir alafenamide
combination in order to reduce the viral load and to increase
CD4+ cell count. Treatment initiation was associated with a better clinical outcome. On September 2021 a new colonoscopy was
performed. The mucosa appeared macroscopically normal from
the rectum to the cecum, no particular lesions were found on the
mucosal surface (Figure 2). The patient was in good general condition. In October 2021 the viral load was significantly decreased
with immune reconstitution of CD4+ T-cells.
Materials and methods
Data sources
The GWAS data was obtained from the MiBioGen consortium, which encompasses a total of 18,340 individuals across 24 distinct cohorts. For genetic data related to AGA, we utilized the GWAS dataset published by EBI in 2018, comprising 66,172 cases and 140,864 control individuals, amounting to a total of 207,036 subjects. Both datasets are derived from populations of European descent and do not differentiate by sex. All data are from publicly available datasets, so no additional ethical approval is required.
Instrumental variable
The instrumental variable in MR studies must satisfy three assumptions: (1) Relevance assumption: There is a robust strong correlation between the instrumental variable and the exposure factor; (2) Independence assumption: The instrumental variables remain independent of confounding factors affecting the “exposure-outcome”; relationship; (3) Exclusivity assumption: Genetic variations influence outcomes solely through exposure factors without any alternative pathways impacting the outcome [12]. SNPs significantly associated with GM (P<1×10-5) were screened while setting (r2<0.001 and kb=10,000) to exclude Linkage Disequilibrium (LD). Using the PhenoScanner database (P<5×10-8; r2>0.8), SNPs directly correlated with confounding factors were excluded based on F-statistic values to remove weak instrumental variables (F<10) [13].
MR analysis
MR analysis primarily employed Inverse Variance Weighting (IVW), Weighted Median, and MR-Egger regression methods [14]. When selected SNPs serve as effective instruments for IV analysis, IVW yields the most precise causal effect estimates [15]. The advantage of Weighted Median lies in its ability to provide consistent causal effect estimates even when some invalid instrumental variables are present [16]. The MR-Egger regression method accounts for potential heterogeneity among instrumental variables and provides adjusted causal effect estimates capable of detecting and correcting pleiotropic bias among instruments [17]. Ultimately, results from IVW will be fitted alongside those from Weighted Median and MR-Egger regression methods [18], with outcomes assessed using Odds Ratios (OR) and corresponding 95% Confidence Intervals (CI) to evaluate potential causal associations.
MR sensitivity analysis
Cochran’s Q test was employed to assess whether there exists significant variability between two sets of results; a P-value less than 0.05 indicates notable differences [19]. To detect horizontal pleiotropy via MR-Egger intercept method: if there is a significant difference between the intercept term and zero along with a P-value for the regression intercept below 0.05, it suggests evidence of horizontal pleiotropy [20]. Additionally, Leave-one-out testing was conducted to ascertain whether individual SNPs exert an influence on MR results.
Results
Instrumental variables
After excluding confounding factors and palindromic SNPs, a total of 69 SNPs were included in the final analysis. The Fstatistics for all selected SNPs in this study exceeded 10, indicating that the results are reliable and there is no concern regarding weak instrument bias. Following the exclusion of potential confounders and palindromic SNPs, we confirmed the inclusion of 69 SNPs for further analysis. The F-statistics associated with these selected SNPs consistently surpassed 10, reinforcing the reliability of our findings while eliminating any possibility of weak instrument bias.
MR analyses
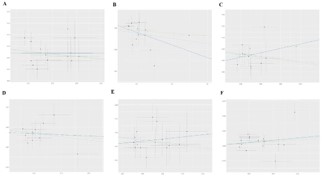
According to the IVW results, increased abundance of certain bacterial taxa was found to be associated with a reduced risk of AGA: the Oxalobacteraceae (OR=0.957, 95% CI 0.9190.996, P=0.033), Paraprevotella (OR=0.953, 95% CI 0.909-0.999, P=0.047), Eubacteriumventriosum group (OR=0.931, 95% CI 0.869-0.997, P=0.043), LachnospiraceaeUCG008 (OR=0.937, 95% CI 0.892-0.985, P=0.011), and Clostridiales (OR=0.917, 95% CI 0.853-0.986, P=0.019) may serve as protective factors against AGA risk. Conversely, an increase in abundance of the Coriobacteriaceae was linked to an elevated risk for AGA (OR=1.103, 95% CI 1.003-1.213, P=0.041), suggesting its potential role as a risk factor for this condition (Table 1). The associations between GM and AGA were revealed through IVW analysis along with MREgger regression and weighted median approaches (Figure 2), where colored lines represent fitting results from these three methodologies.
Table 1: MR analysis results.
| Conclusion |
Exposure |
Method |
OR (CI95%) |
β |
SE |
P |
| Androgenetic alopecia |
Oxalobacteraceae |
IVW |
0.957 (0.9190.996) |
-0.043 |
0.02 |
0.033 |
| Weighted Median |
0.955 (0.9061.006) |
-0.045 |
0.026 |
0.045 |
| MR-Egger |
0.999 (0.8481.177) |
-0.001 |
0.083 |
0.099 |
| Paraprevotella |
IVW |
0.953 (0.9090.999) |
-0.047 |
0.024 |
0.047 |
| Weighted Median |
0.976 (0.9151.041) |
-0.023 |
0.032 |
0.047 |
| MR-Egger |
0.888 (0.7581.042) |
-0.117 |
0.081 |
0.174 |
| Eubacterium ventriosum group |
IVW |
0.931 (0.8690.997) |
-0.070 |
0.035 |
0.043 |
| Weighted Median |
0.924 (0.841.017) |
-0.078 |
0.048 |
0.011 |
| MR-Egger |
1.136 (0.8451.527) |
0.128 |
0.150 |
0.413 |
| LachnospiraceaeUCG008 |
IVW |
0.937 (0.8920.985) |
-0.064 |
0.025 |
0.011 |
| Weighted Median |
0.949 (0.8871.015) |
-0.051 |
0.034 |
0.013 |
| MR-Egger |
0.977 (0.7591.259) |
-0.022 |
0.128 |
0.866 |
| Clostridiales |
IVW |
0.917 (0.853-0.986) |
-0.085 |
0.036 |
0.019 |
| Weighted Median |
0.909 (0.8261.001) |
-0.094 |
0.048 |
0.032 |
| MR-Egger |
1.079 (0.7681.516) |
0.076 |
0.173 |
0.665 |
| Coriobacteriaceae |
IVW |
1.103 (1.0031.213) |
0.098 |
0.048 |
0.041 |
| Weighted Median |
1.082 (0.9751.201) |
0.079 |
0.052 |
0.042 |
| MR-Egger |
1.135 (0.7691.675) |
0.127 |
0.198 |
0.533 |
MR sensitivity analyses
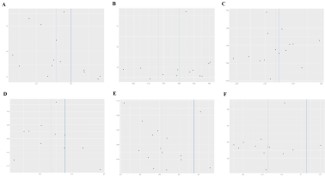
Cochran’s Q test assessed heterogeneity within data results from both IVW and MR-Egger regression analyses; no evidence of heterogeneity was detected (P>0.05). To evaluate horizontal pleiotropy, we examined whether there was deviation between the intercept term obtained from MR-Egger regression and zero; our findings indicated absence thereof (P>0.05) (Table 2). Funnel plot analysis demonstrated that all SNP distributions were approximately symmetrical, confirming robustness in causal associations. Leave-one-out tests evaluated each individual SNP’s impact on overall causal relationships; upon removing one SNP at a time, remaining ones consistently fell on one side relative to null effect line without identifying any single influential outlier which suggests that our MR analytical outcomes are not driven by any particular SNP but rather exhibit substantial robustness across analyses performed. By assessing distances between intercept terms derived from MR-Egger regressions compared against zero points enables us ascertain presence or absence concerning horizontal pleiotropy; analyses yielded conclusive evidence negating such effects (P>0.05) (Table 2). Furthermore, funnel plots illustrated near-symmetrical distribution among all considered variants affirming stability inherent within observed causal links (Figure 3). Lastly leave-one-out assessments gauged influence exerted by every variant upon collective causation revealing consistent positioning beyond threshold delineating non-effectiveness post removal thus underscoring resilience exhibited throughout conducted investigations (Figure 4) thereby validating integrity surrounding resultant conclusions drawn herein.
Table 2 Results of sensitivity analysis.
| Microbiota |
Cochran's Q test |
MR-Egger regression |
| Q |
P |
Egger-intercept |
P |
| Oxalobacteraceae |
14.545 |
0.336 |
-0.005 |
0.605 |
| Paraprevotella |
13.387 |
0.341 |
0.007 |
0.385 |
| Eubacteriumventriosum group |
11.473 |
0.488 |
-0.014 |
0.202 |
| LachnospiraceaeUCG008 |
8.417 |
0.588 |
-0.004 |
0.748 |
| Clostridiales |
9.738 |
0.781 |
-0.011 |
0.354 |
| Coriobacteriaceae |
25.522 |
0.029 |
-0.001 |
0.885 |
Discussion
This study analyzes the relationship between GM and AGA using MR methods. The findings indicate that five specific GM groups may serve as protective factors against AGA: Oxalobacteraceae (OR=0.957, 95% CI 0.919-0.996, P=0.033), Paraprevotella (OR=0.953, 95% CI 0.909-0.999, P=0.047), Eubacteriumventriosum group (OR=0.931, 95% CI 0.869-0.997, P=0.043), LachnospiraceaeUCG008 (OR=0.937, 95% CI 0.892-0.985, P=0.011), and Clostridiales (OR=0.917, 95% CI 0.853-0.986, P=0.019). Conversely, the Coriobacteriaceae (OR=1.103, 95% CI 1.003-1.213, P=0.041) may be a potential risk factor for AGA. The research suggests a bidirectional association between GM and AGA; specifically, levels of the enzyme 5α-reductase in hair follicles from balding areas are significantly higher than those in non-balding regions [21]. This enzyme facilitates the conversion of testosterone to dihydrotestosterone within tissues-leading to follicular miniaturization and subsequent abnormal hair growth patterns over time which can result in gradual follicle atrophy until complete loss occurs [22]. The gastrointestinal tract is one of the largest epithelial tissues in the human body with a surface area ranging from approximately 250 to 400 square meters housing over 500 species of bacteria. The predominant bacterial phyla include Bacteroidetes and Firmicutes while Proteobacteria Ac-tinobacteria and Fusobacteriaphyla are commonly found in these areas [23]. Rebello D. conducted an experiment wherein they introduced Clostridium difficile into patients’ intestines via fecal microbiota transplantation techniques, resulting in observable hair regrowth on various parts including scalp, facial, and armregions [24]. Additionally, Alkhalifah A. analyzed stool samples from patients with alopecia areata constructing predictive models for disease status. They discovered that models based on Subdoligranulum and Butyricicoccus could accurately predict patients conditions [25]. Moreover, the alteration of GM can interact with immune systems through harmful metabolites increasing pro-inflammatory cells, cytokines, and metabolites circulating into distal organs such as brain, lungs, and skin leading to inflammation states within specific extraintestinal organs [26]. The onset of skin diseases not only triggers changes within cutaneous microbiomes but also induces corresponding modifications among intestinal flora [27], suggesting possible bidirectional signaling pathways between skin and gut. This pathway plays a crucial role in alleviating inflammation across extraintestinal organs while improving symptom atology associated with disease [28]. Liu C. identified alterations in GM structure among patients suffering from melasma as well as the biological effects offirmicutes with in microbial communities potentially influencing β-glucuronidase production alongside androgen synthesis or metabolism thereby contributing to melasma development [29]. In summary, these studies reveal a potential association between Clostridium, Methanobrevibacter, Rhodococcus, Oxalobacter, and Faecalibacterium with neuro inflammation and immune damage in AGA.
The study reveals associations between lifelong carriers of alleles related to AGA in the general population, distinguishing it from traditional randomized controlled trials. This allows for a comprehensive assessment of the role of GM in the prevention or treatment of AGA [30]. MR analysis mitigates interference from confounding factors and bidirectional causal relationships in association analyses while considering the long latency period of diseases and the impact of individuals’ lifetime exposure to risk factors, significantly reducing inherent biases present in observational studies. Causal effects were evaluated using inverse variance weighting, weighted median methods, and MREgger regression analysis, with sensitivity analyses further confirming the stability and reliability of results. The data for this MR study was sourced from populations of European descent, utilizing recent GWAS findings on both GM and AGA, which provided larger sample sizes and higher statistical power. However, this research has certain limitations. Firstly, GWAS data lacks detailed statistics regarding participant sex ratios, environmental factors, racial differences, and disease duration; thus, complicating comparisons of causal effects across different subgroups. The presence of multiple etiologies and subtypes may also affect the precision of research outcomes. Secondly, this study exclusively included individuals from European ancestry without analyzing other ethnic groups; although this helps reduce population stratification bias, its applicability to other racial groups may be limited [31]. While this study confirms that GM has a protective effect against AGA, it does not explore potential mediating factors influencing these results. Interventions solely targeting GM may not provide optimal solutions for mitigating or treating AGA risk due to numerous other non-genetic factors, such as unemployment status, psychosocial influences, poverty levels, educational opportunities and access to healthcare services, which were not incorporated into this research nor obtainable through GWAS methodologies.
Conclusion
This study provides evidence for a potential causal relationship between gut microbiota composition and AGA. Dysbiosis may influence AGA through immune modulation, androgen metabolism, and systemic pathways. These findings highlight the importance of gut health in AGA pathogenesis and open new avenues for therapeutic intervention. Future research should explore the underlying mechanisms and evaluate the efficacy of microbiota-targeted therapies in preventing or treating AGA, offering a novel perspective on managing this widespread condition.
Declarations
Acknowledgments: The authors thank all participants and investigators who provided the GWAS data.
Author contributions: Shuning Liu: Conceptualization, Formal analysis, Methodology, Software, Project administration, Investigation, Data curation, Resources, Supervision, Validation, Writing - original draft, Writing - review & editing.
Debin Xu: Project administration, Supervision, Writing - review & editing.
Institutional review board statement: All data are publicly available GWAS datasets, therefore no additional ethical approval was required.
Conflicts of interest: The authors declare no conflicts of interest.
References
- Maryam N, Amir T, Hasan S, et al. Distinguishing immunohistochemical features of alopecia areata from androgenic alopecia. Journal of Cosmetic Dermatology. 2018; 18: 422-426.
- Sorbellini E, Pinto D, Marzani B, et al. Drug treatment for androgenetic alopecia: first Italian questionnaire survey on what dermatologists think about finasteride. Dermatology and Therapy. 2018; 8: 259-267.
- Kaliyadan F, Nambiar A, Vijayaraghavan S. Androgenetic alopecia: an update. Indian Journal of Dermatology, Venereology and Leprology. 2013; 79: 613-625.
- Hammes SR, Levin ER. Impact of estrogens in males and androgens in females. J Clin Invest. 2019; 129: 1818-1826.
- Colucci R, Moretti S. Implication of Human Bacterial Gut Microbiota on Immune-Mediated and Autoimmune Dermatological Diseases and Their Comorbidities: A Narrative Review. Dermatol Ther (Heidelb). 2021; 11: 363-384.
- Pascal M, Perez-Gordo M, Caballero T, et al. Microbiome and allergic diseases. Front Immunol. 2018; 9: 1584.
- Salem I, Ramser A, Isham N, et al. The gut microbiome as a major regulator of the gut-skin axis. Front Microbiol 2018; 9: 1459.
- Kim So, Sheikh Hi, Ha Sd, et al. G-CSF-mediated inhibition of JNK is a key mechanism for Lactobacillus rhamnosus-induced suppression of TNF production in macrophages. Cellular Microbiology. 2006; 8: 1958-1971.
- Wang K, Han SZ. Effect of selection bias on two sample summary data based Mendelian randomization. Scientific Reports. 2021; 11: 7585.
- Burgess S, Small DS, Thompson SG. A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res. 2017; 26: 2333-2355.
- Emdin CA, Khera AV, Kathiresan S. Mendelian randomization. JAMA. 2017; 318: 1925.
- Bowden J, Davey Smith G, Haycock PC, et al. Consistent estimation in mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol. 2016; 40: 304-314.
- Lu JP, Lu Y, Yang H, et al. Characteristics of high cardiovascular risk in 1.7 million Chinese adults. Annals of Internal Medicine. 2019; 170: 298-308.
- Shen J, Zhou H, Liu J, et al. A modifiable risk factors atlas of lung cancer: A Mendelian randomization study. Cancer Med. 2021; 10: 4587-4603.
- Burgess S, Bowden J, Fall T, et al. Sensitivity analyses for robust causal inference from Mendelian randomization analyses with multiple genetic variants. Epidemiology. 2017; 28: 30-42.
- Slob EAW, Groenen PJF, Thurik AR, et al. A note on the use of egger regression in mendelian randomization studies. Int J Epidemiol. 2017; 46: 2094-2097.
- Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol. 2015 ; 44: 512-525.
- Yuan J, Wang D, Zhang Y, et al. Genetically predicted obesity and risk of hip osteoarthritis. Eat Weight Disord. 2023; 28: 11.
- Greco M FD, Minelli C, Sheehan NA, et al. Detecting pleiotropy in Mendelian randomization studies with summary data and a continuous outcome. Stat Med. 2015; 34: 2926-2940.
- Gala H, Tomlinson I. The use of Mendelian randomization to identify causal cancer risk factors: promise and limitations. J Pathol. 2020; 250: 541-554.
- York K, Meah N, Bhoyrul B, Sinclair R. A Review of The Treatment of Male Pattern Hair Loss. Expert Opinion on Pharmacotherapy. 2020; 21: 603-612.
- Hammes Sr, Levin ER. Impact of estrogens in males and androgens in females. The Journal of Clinical Investigation. 2019; 129: 1818-1826.
- Thursby E, Juge N. Introduction to the human gut microbiota. Biochem J. 2017; 474: 1823-1836.
- Rebello D, Wang E, Yen E, et al. Hair Growth in Two Alopecia Patients after Fecal Microbiota Transplant. ACG Case Rep J. 2017; 4: e107.
- Moreno-Arrones OM, Serrano-Villar S, Perez-Brocal V, et al. Analysis of the gut microbiota in alopecia areata: identification of bacterial biomarkers. J Eur Acad Dermatol Venereol. 2020; 34: 400-405.
- Levy M, Kolodziejczyk A. A, Thaiss C. A, et al. Dysbiosis and the immune system. Nat Rev Immunol. 2017; 17: 219-232.
- O’Neill CA, Monteleone G, McLaughlin JT, et al. The gut-skin axis in health and disease: A paradigm with therapeutic implications. BioEssays. 2016; 38: 1167-1176.
- Salem I, Ramser A, Isham N, et al. The Gut Microbiome as a Major Regulator of the Gut-Skin Axis. Front Microbiol. 2018; 9: 1459.
- Liu C, He D, Yu A, et al. Correlation analysis between gut microbiota characteristics and melasma. Front Microbiol. 2022; 13: 1051653.
- Smith GD, Ebrahim S. Mendelian randomization: can genetic epidemiology contribute to understanding environmental determinants of disease. Int J Epidemiol. 2013; 32: 1-22.
- Bowden J, Holmes MV. Meta-analysis and mendelian randomization: a review. Res Synth Methods. 2019; 10: 486-49.